I’m co-organiser of this slightly-different in October 2023 at the Forschungszentrum Jülich, Germany. Rather than following the traditional format of 3-4 day populated by talks with the odd poster session, this is an extended workshop made up of six mini-workshops. Since it is focussed on python-based tools for biomolecular simulations, of which there are an increasing number, the first mini-workshop will be a bootcamp that I will be lead instructor on (helped by David Dotson from ASU). I’m also leading the next mini-workshop on analysing biomolecular simulation data.
molecular dynamics
New Publication: Alchembed
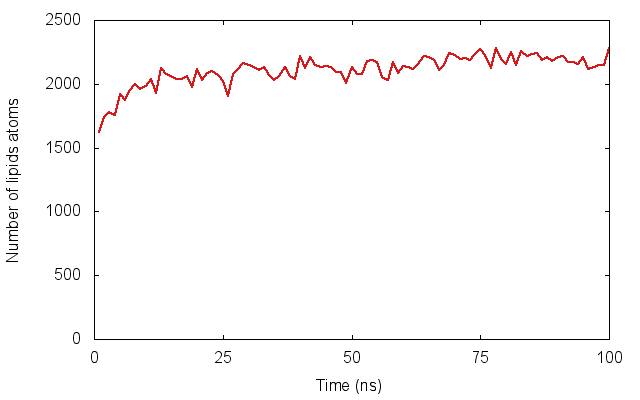
In much of my research I’ve looked at how proteins embedded in cell membranes behave. An important part in any simulation of a membrane protein is, obviously, putting it into a model membrane, often a square patch of several hundred lipid molecules. This is surprisingly difficult: although a slew of methods have been published, none of them can embed several proteins simultaneously into a complex (non-flat) arrangement of lipids. For example, a virus, as shown in our recent paper.
Here we introduce a new method, dubbed Alchembed, that uses an alternative way, borrowed from free energy calculations, of “turning on�? the van der Waals interactions between the protein and the rest of the system. We show how it can be used to embed five different proteins into a model vesicle on a standard workstation. If you want to try it out, there is a on GitHub. This assumes you have is setup
You can get the paper for free from .
Installing GROMACS with MPI support on a Mac
GROMACS is an optimised molecular dynamics code, primarily used for simulating the behaviour of proteins. To compile GROMACS you need, well, some compilers. I install gcc using . Note that this requires you to . Then it is easy to install gcc version 4.9 by
sudo port install gcc49
(and yes, I know about , but I still find has more of the things I want than brew). So, once you’ve done a bit of preparation, on a Mac is easy. Once you’ve downloaded the source code tar ball.
tar xvf gromacs-5.0.2.tar.gz
cd gromacs-5.0.2
mkdir build
cmake .. -DGMX_BUILD_OWN_FFTW=ON -DCMAKE_INSTALL_PREFIX='/usr/local/gromacs/5.0.2/‘
make
sudo make install
Note that this will install it in /usr/local/gromacs/5.0.2 so you can keep multiple versions on the same machine and swap between them in a sane way by sourcing the GMRXC file, for example
source /usr/local/gromacs/4.6.7/bin/GMXRC
Adding MPI support on a Mac is trickier. This appears mainly to be because the gcc compilers from MacPorts (or clang from Xcode) don’t appear to support OpenMPI. You will know because when you run the cmake command you get a load of failures starting about ten lines down, such as
-- Performing Test OpenMP_FLAG_DETECTED - Failure
I managed to get a working version using the following approach; it is likely there are better (if you know, please leave a comment), but it has the virtue of working. First we need to install OpenMPI.
sudo port install openmpi
Now we need a compiler that supports OpenMPI. If you dig around in the MacPorts tree you can find some.
sudo port install openmpi-devel-gcc49
Finally, we can follow the steps above (I just mkdir build-mpi subfolder in the above source folder and then cd to it), but now we need a (slightly) complex cmake instruction
cmake .. -DGMX_BUILD_OWN_FFTW=ON
-DGMX_BUILD_MDRUN_ONLY=on
-DCMAKE_INSTALL_PREFIX=/usr/local/gromacs/5.0.2
-DGMX_MPI=ON -DCMAKE_C_COMPILER=mpicc-openmpi-devel-gcc49
-DCMAKE_CXX_COMPILER=mpicxx-openmpi-devel-gcc49
-DGMX_SIMD=SSE4.1
This is only going to build an MPI version of mdrun (which makes sense) and will install mdrun_mpi alongside the regular compiled binaries we did first. We have to tell cmake what all the new fancy compilers are called and, unfortunately, these don’t support AVX SIMD instructions so we have to fall back to SSE4.1. Experience suggests this doesn’t impact performance as much as you might think. Now you can run things like on your workstation!
A simple tutorial on analysing membrane protein simulations.
I’m teaching on how to analyse membrane protein simulations next week at the University of Bristol as part of a arranged by . As it is only 90 minutes long, it only covers two simple tasks but I show how you can do both with (a python module) or in Tcl in . Rather than write something and just distribute it to the people who are coming to the course, I’ve put the whole tutorial, including trajectory files and all the example code . Please feel free to clone it, make changes and send a pull request (or just send me any comments).
GROMACS 4.6: Running on GPUs
I mentioned before that I would write something on running on GPUs. Let’s imagine we want to simulate a solvated lipid bilayer containing 6,000 lipids for 5 µs. The total number of is around 137,000 and the box dimensions are roughly 42x42x11 nm. Although this is smaller than the benchmark we looked at last time, it is still a challenge to run on a workstation. To see this let’s consider running it on my MacPro using GROMACS 4.6.1. The machine is an and has 2 Intel Xeons, each with 4 cores. Using 8 MPI processes gets me 132 ns/day, so I would have to wait 38 days for 5 µs. Too slow!
You have to be careful installing non-Apple supported NVidia GPUs into MacPros, not least because you are limited to 2x6pin power connectors. Taking all this into account, the best I can do without doing something drastic to the power supply is to install an . Since I only have one GPU, I can only run one MPI process, but this can spawn multiple OpenMP threads. For this particular system, I get the best performance with 4 threads (134 ns/day) which is the same performance I get using all 8 cores without the GPU. So when I am using just a single core, adding in the GPU increases the performance by a factor of 3.3x. But as I add additional cores, the increase afforded by the single GPU drops until the performance is about the same at 8 cores.
Now let’s try something bigger. Our lab has a small Intel (Sandy Bridge) computing cluster. Each node has 12 cores, and there are 8 nodes, yielding a maximum of 96 cores. Running on the whole cluster would reduce the time down to 6 days, which is a lot better but not very fair on everyone else in the lab. We could try and get access to Tier-1 or Tier-0 supercomputers but, for this system, that is overkill. Instead let’s look at a Tier-2 machine that uses GPUs to accelerate the calculations.
The University of Oxford, through , has access to the machines owned by the Centre for Innovation. One of these, , is a GPU-based cluster. We shall look at one of the partitions; this has 60 nodes, each with two 6-core Intel processors and 3 NVIDIA M2090 Tesla GPUs. For comparison, let’s run without the GPUs. The data shown are for simulations with only 1 OpenMP thread per MPI process. So now let’s run using the GPUs (which is the point of this cluster!). Again just using asingle OpenMP thread per MPI process/GPU (shown on graph) we again find a performance increase of 3-4x. Since there are 3 GPUs per node, and each node has 12 cores, we could run 3 MPI processes (each attached to a GPU) on each node and each process could spawn 1, 2, 3 or 4 OpenMP threads. This uses more cores, but since they probably would be sitting idle, this is a more efficient use of the compute resource. Trying 2 or 3 OpenMP threads per MPI process/GPU lets us reach a maximum performance of 1.77 µs per day, so we would get our 5 µs in less than 3 days. Comparing back to our cluster, we can get the same performance of our 96-core local cluster using a total of 9 GPUs and 18 cores on EMERALD.
Finally, let’s compare EMERALD to the Tier-1 PRACE supercomputer CURIE. CURIE was the . For this comparison we will need to use a bigger benchmark, so let’s us . It has 9x the number of lipids, but because I had to add extra water ends up being about 15x bigger at 2.1 million particles. Using 24 GPUs and 72 cores, EMERALD manages 130 ns/day. To get the same performance on CURIE requires 150 cores and ultimately CURIE tops out at 1,500 ns/day on 4,196 cores. Still, EMERALD is respectable and shows how it can serve as a useful bridge to Tier-1 and Tier-0 supercomputers. Interestingly, CURIE also has a “hybrid�? partition that contains 144 nodes, each with 2 Intel Westmere processors and 2 NVIDIA M2090 Tesler GPUs. I was able to run on up to 128 GPUs with 4 OpenMP threads per MPI/GPU, making a total of 512 cores. This demonstrates that GROMACS can run on large numbers of GPU/CPUs and that such hybrid architectures are viable as supercomputers (for GROMACS at least).
GROMACS 4.6: Scaling of a very large coarse-grained system
So if I have a particular system I want to simulate, how many processing cores can I harness to run a single version 4.6 job? If I only use a few then the simulation will take a long time to finish, if I use too many the cores will end up waiting for communications from other cores and so the simulation will be inefficient (and also take a long time to finish). In between is a regime where the code, in this case GROMACS, scales well. Ideally, of course, you’d like linear scaling i.e. if I run on 100 cores in parallel it is 100x faster than if I ran on just one.
The rule of thumb for GROMACS 4.6 is that it scales well until there are only ~130 atoms/core. In other words, the more atoms or beads in your system, the larger the number of computing cores you can run on before the scaling performance starts to degrade.
As you might imagine there is a hierarchy of computers we can run our simulations on; this starts at humble workstations, passes through departmental, university and regional computing clusters before ending up at national (Tier 1) and international (Tier 0) high performance computers (HPC).
In our lab we applied for, and , a set of European Tier 0 supercomputers through PRACE. These are currently amongst the . We tested five supercomputers in all: (Paris, France; green lines), (Barcelona, Spain; black line), (Bologna, Italy; lilac), (Munich, Germany; blue) and (Stuttgart, Germany; red ). Each has a different architecture and inevitably some are slightly newer than others. CURIE has three different partitions, called thin, fat and hybrid. The thin nodes constitute the bulk of the system; the fat nodes have more cores per node whilst the hybrid nodes combine conventional CPUs with GPUs.
We tested a coarse-grained 54,000 lipid bilayer (2.1 million beads) on all seven different architectures and the performance is shown in the graph – note that the axes are logarithmic. Some machines did better than others; FERMI, which is an , appears not to be well-suited to our benchmark system, but then one doesn’t expect fast per-core performance on a BlueGene as that is not how they are designed. Of the others, MareNostrum was fastest for small numbers of cores, but its performance began to suffer if more than 256 cores were used. SuperMUC and the Curie thin nodes were the fastest conventional supercomputers, with the Curie thin nodes performing better at large core counts. Interestingly, the Curie hybrid GPU nodes were very fast, especially bearing in mind the CPUs on these nodes are older and slower than those in the thin nodes. One innovation introduced into GROMACS 4.6 that I haven’t discussed previously is one can now run either using purely MPI processes or a . We were somewhat surprised to find, that, in nearly all cases, the pure MPI approach remained slightly faster than the new hybrid parallelisation.
Of course, you may see very different performance using your system with GROMACS 4.6. You just have to try and see what you get! In the next post I will show some detailed results on using GROMACS on GPUs.
GROMACS 4.6
is a scientific code designed to simulate the dynamics of small boxes of stuff, that usually contain a protein, water, perhaps a lipid bilayer and a range of other molecules depending on the study. It assumes that all the atoms can be represented as points with a mass and an electrical charge and that all the bonds can modelled using simple harmonic springs. There are some other terms that describe the bending and twisting of molecules and all of these, when combined with two long range terms, which take into account the repulsion and attraction between electrical charges, allow you to calculate the force on any atom due to the positions of all the other atoms. Once you know the force, you can calculate where the atom will be a short time later (often 2 fs) but of course the positions have changed so you have to recalculate the forces. And so on.
Anyway, I use GROMACS a lot in my research and the most recent major version, 4.6, . In this post I’m going to briefly describe my experience with some of the improvements. First off, so much has changed that I think it would have been more accurate to call this GROMACS 5.0. For example, version 4.6 is a lot faster than version 4.5. I typically use three different benchmarks when measuring the performance; one is an all-atom simulation of a bacterial peptide transporter in a lipid bilayer (78,033 atoms). The other two are both coarse-grained models of a lipid bilayer using the – the difference is one has 6,000 lipids (137,232 beads), the other 54,000 (2,107,010 beads). Ok, so how much faster is version 4.6? It is important here to bear in mind that GROMACS was already very fast since a lot of effort had been put into optimising the loops that the code spends most of its time running. Even so, version 4.6 is between 20-120% faster when using either of the first two benchmarks, and in some cases even faster. How? Well, it seems the developers using commands. One important consequence of this is that it is and, since you have to specify which SIMD instruction sets to use, you may need several different versions of the key binary, mdrun. For example, you may want a version compiled using AVX SIMD instruction sets for recent CPUs, but also a version compiled using an older SSE SIMD instruction set. The latter will run on newer architectures, but it will be slower. You must never run a version compiled with no SIMD instruction sets as this can be 10x slower!
The other big performance improvement is that GROMACS 4.6 now uses GPUs seamlessly. The calculations are shared between any GPUs and the CPUs and GROMACS will even shift the load to try and share it equally. , one of the GROMACS developers, gave an on this subject in April 2013. A GPU here just means a reasonable consumer graphics card, such as an NVIDIA GTX680, that has compute capability of 2.0 or higher. So, how much performance boost do we see? I typically see a boost of 2.1-2.7x for the atomistic benchmark and 1.4-2.2x for the first, smaller coarse-grained benchmark. Just for fun, you can try running a version of GROMACS compiled with no SIMD instructions with a GPU (and without a GPU) and then you can get a performance increase of 10x.
Before I finish, I was given some good advice on running GROMACS benchmarks. Firstly, make sure you use the -noconfout mdrun option since this prevents it from writing a final .gro file as this takes some time. Secondly setup a .tpr file that will run for a long (wallclock) time even on a large number of cores and then use the -resethway option in combination with a time limit, such as -maxh 0.25, as this would then reset the timers after 7.5 min and record how many steps were calculated between 7.5 and 15 minutes. From experience a bit of time spent writing some good BASH scripts to automatically setup, run and analyse the benchmarking simulations really pays off in the long run.
In future posts I’ll talk about the scaling of GROMACS 4.6 (that is where the third benchmark comes in) and also look at the GPU performance in a bit more detail.